AtaGenix Laboratories
AtaGenix Laboratories
AtaGenix offers yeast recombinant protein expression using Pichia pastoris as the primary host, with Saccharomyces cerevisiae available for GRAS-requirement applications. Yeast combines eukaryotic processing (glycosylation, disulfide bonds, proper folding) with the ease of microbial culture — enabling high-density fermentation and efficient secreted expression at a fraction of mammalian cell costs.
Pichia
Primary Host
6–10 wks
Typical Timeline
5 Vectors
Expression Options
Scalable
Flask to Fermenter
The Pichia system is particularly well-suited for secreted proteins — the host secretes very few native proteins, making purification from culture supernatant straightforward. Multiple strains (GS115, KM71, X33) and vectors (pPICZα, pPIC9K for secretion; pAO815, pPIC3K, pPIC3.5K for intracellular) are available with a library of 5 signal peptides for optimized secretion.
| Expression Mode | Strains | Vectors |
| Secreted | GS115 (Mut+), KM71 (Muts), X33 | pPICZα A/B/C, pPIC9K |
| Intracellular | GS115 (Mut+), KM71 (Muts) | pAO815, pPIC3K, pPIC3.5K |
Signal peptides for secretion: AlphaMF, Dse4, Exg1, Scw11, SAlb. All vectors use the AOX1 methanol-inducible promoter.
High-Throughput Screening
Not every clone expresses equally. Our high-throughput screening platform rapidly identifies top-expressing clones under optimized induction conditions before committing to scale-up — saving weeks of trial and error.
Clean Secretion = Simple Purification
Pichia secretes very few native proteins into the medium. Your target protein dominates the supernatant, reducing purification steps and boosting recovery compared to intracellular E. coli workflows.
Eukaryotic PTMs at Microbial Cost
Glycosylation, disulfide bonds, phosphorylation, and lipid modifications — all available in a system that grows to high cell density on inexpensive glycerol/methanol media. A fraction of CHO/HEK293 cost.
AtaGenix provides a complete Pichia pastoris and S. cerevisiae expression platform with strains GS115, KM71, and X33, five expression vectors (pPICZα, pPIC9K, pAO815, pPIC3K, pPIC3.5K), and a library of 5 signal peptides for secretion optimization. Our high-throughput screening platform identifies top-expressing clones before scale-up, and large-scale fermentation capability supports projects from mg to gram-level production. Typical timeline: 6–10 weeks from gene sequence to delivered protein.
Service Scope
| ✓ Gene synthesis & codon optimization for yeast | ✓ Yeast expression vector construction |
| ✓ Pichia: GS115, KM71, X33 hosts | ✓ AOX1 methanol-inducible promoter system |
| ✓ 5 signal peptides for secretion optimization | ✓ High-throughput clone screening |
| ✓ Flask to fermenter scale-up | ✓ Chromatography-based purification |
| ✓ QC: SDS-PAGE, Western Blot + detailed technical report | |
| Target Type | Why Yeast |
| Secreted proteins | Low host protein background simplifies purification — fewer contaminants, fewer steps |
| Industrial enzymes | High-density fermentation on inexpensive media enables gram-level production at low cost |
| Glycoproteins (simple glycans) | Eukaryotic glycosylation and disulfide bond formation without mammalian cost |
| Food-grade & diagnostic proteins | S. cerevisiae has GRAS status; Pichia offers low-cost, high-yield secretion for diagnostic antigens |
Pichia
Primary Host
Secreted
Expression Mode
g/L
Yield Potential
Scalable
Flask to Fermenter
Every yeast project follows a standardized pipeline from gene design through fermentation to purified protein. Clone screening via our high-throughput platform is the critical optimization step. Total timeline: 6–10 weeks.
01
Gene & Vector
2–4 weeks
Codon optimization
Gene synthesis
Subcloning (1 week)
02
Pilot Expression
2–3 weeks
Transformation & clone screening
High-throughput optimization
SDS-PAGE/WB verification
03
Scale-Up & Purify
1–2 weeks
Fermentation under optimized conditions
Chromatography purification
Concentration & buffer exchange
04
QC & Delivery
Included
SDS-PAGE, Western Blot
Technical report
Protein sample delivery
Step 1 — Gene & Vector (2–4 weeks): Codon optimization for Pichia expression, gene synthesis, and subcloning into the appropriate vector (pPICZα A/B/C or pPIC9K for secretion; pAO815, pPIC3K, or pPIC3.5K for intracellular). Signal peptide selected from AlphaMF, Dse4, Exg1, Scw11, or SAlb. Skipped if client provides a ready-to-use construct.
Step 2 — Pilot Expression (2–3 weeks): Transformation into Pichia strains (GS115, KM71, or X33), high-throughput clone screening to identify top expressers, and optimization of induction conditions (methanol concentration, temperature, time). SDS-PAGE and Western Blot confirm expression before scale-up.
Step 3 — Scale-Up & Purification (1–2 weeks): Large-scale fermentation under optimized conditions. Chromatography-based purification workflow tailored to your protein. Concentration and buffer exchange to your specifications.
Step 4 — QC & Delivery: SDS-PAGE and Western Blot QC. Protein delivered with a detailed technical report including expression conditions, purification methods, and QC data.
Yeast sits between E. coli and mammalian cells in terms of capability and cost. Choose yeast when your protein needs eukaryotic folding and simple glycosylation, but mammalian costs are not justified.
| Feature | E. coli | Yeast | Insect | Mammalian |
|---|---|---|---|---|
| Glycosylation | None | High-mannose | Paucimannose | Complex / human |
| Secretion | Limited | Excellent | Good | Good |
| Scalability | High | Very high | Moderate | Moderate |
| Relative Cost | $ | $–$$ | $$ | $$$ |
Need a different system? E. coli for lowest cost non-glycosylated targets. HEK293 or CHO for human-type complex glycosylation. Insect cells for complex multi-subunit proteins.
What is the difference between Pichia pastoris and Saccharomyces cerevisiae?
Pichia pastoris is the preferred choice for secreted protein production — it grows to very high cell densities under methanol induction (AOX1 promoter) and secretes very few native proteins, making purification straightforward. S. cerevisiae is a GRAS organism with well-established genetics, suitable for food-grade and regulatory-friendly applications. AtaGenix's primary yeast platform is Pichia; S. cerevisiae is available for specific applications — consult our team for recommendations.
How long does yeast protein expression take?
A standard project takes 6–10 weeks: gene synthesis and codon optimization (2–4 weeks), subcloning (1 week), expression testing and high-throughput clone screening (2–3 weeks), and fermentation with purification (1–2 weeks). Timeline can be shortened if you provide a ready-to-use expression construct.
When should I choose yeast over E. coli or mammalian cells?
Choose yeast when your protein needs eukaryotic folding, disulfide bonds, or simple (high-mannose) glycosylation, but mammalian costs are not justified. Yeast is especially strong for secreted proteins, industrial enzymes (high-density fermentation at low cost), and GRAS-requirement applications. For non-glycosylated targets, E. coli is faster and cheaper. For complex or human-type glycosylation, choose CHO or HEK293.
What QC is included?
Every project includes SDS-PAGE (purity) and Western Blot (identity), delivered with a detailed technical report covering expression conditions, purification methods, and QC results. Protein sample is provided in your specified format. Additional characterization is available upon request.
Yeast glycosylation produces high-mannose type glycans which differ from mammalian complex glycans. Timelines and yields are target-dependent. Quote-based pricing.
Explore real-world yeast protein expression case studies from AtaGenix. Our Pichia pastoris and Saccharomyces cerevisiae platforms deliver high-yield recombinant proteins with proper folding and glycosylation — supporting enzyme production, diagnostic antigen development, and antibody screening applications.
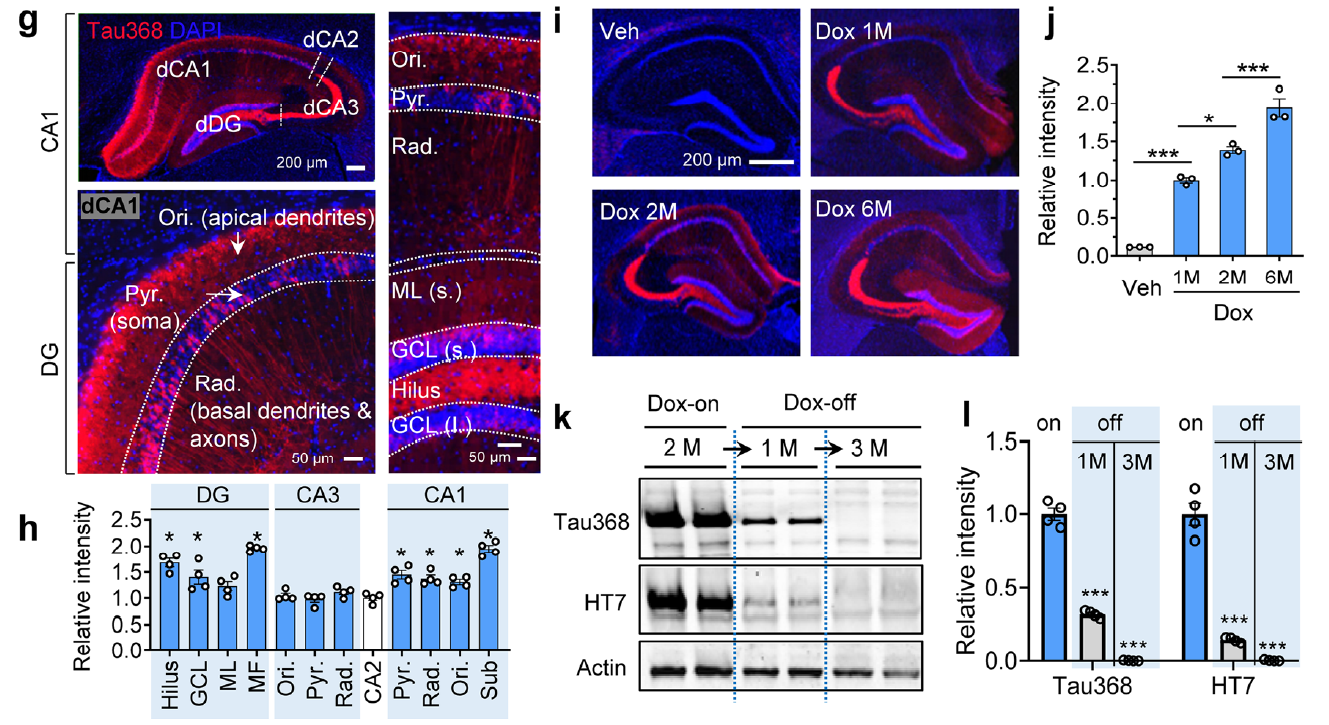
AtaGenix provides highly specific anti-tauN368 monoclonal antibodies to support Alzheimer’s disease research using the hTau368 transgenic mouse model. These antibodies enable precise detection of truncated tau fragments in Western blot and immunofluorescence experiments, validating tau accumulation, phosphorylation, and associated cognitive deficits in the hippocampus. With exceptional specificity and stability, AtaGenix’s solutions empower researchers to explore tau pathology mechanisms and advance tau-targeted therapeutic development.